If you think of the human body as consisting of a team of organs that plays against a defense of diseases, then the AT1 receptor is a key player — especially when it comes to attacks on the heart and kidneys. To better understand this protein’s role, researchers from Oregon State University and the University of Vermont used Expanse at the San Diego Supercomputer Center (SDSC) at UC San Diego and Anton 2 at the Pittsburgh Supercomputing Center (PSC) to show the effect of physical stimuli on the structural properties of the AT1 receptor.
The angiotensin II type 1 (AT1) receptor is heavily studied within the context of activation/inactivation by drugs, but the team’s newly run simulations have revealed that the protein can also be directly activated by mechanical forces through stimuli such as external pressure or shear acting on the surface of the cells. These findings have been published in Nature Communications.
“The AT1 receptor is a very important drug target due to its essential function in many diseases,” Associate Professor of Biochemistry at Oregon State University Juan Vanegas, who directed the project, said. “Understanding how physical stimuli affect its function not only helps us understand the basic biology of this protein, but it may help the scientific community to develop better treatments for diseases associated with this receptor.”
G-protein coupled receptors (GPCRs), including the AT1 receptor, can sometimes malfunction. This makes them some of the biggest targets for drug discovery. Specifically, physicians prescribe medicine for people facing diseases such as hypertension and kidney disease with the prescription targeting this well-known AT1 receptor, which is a membrane protein essential for heart and kidney function.
“AT1 receptors are known to be mechanically sensitive (i.e., their function is modulated by external forces in addition to chemical activation), but this mechanical activation has not been widely studied,” Vanegas explained. “We first used Expanse at SDSC to create very long all-atom simulations and free-energy calculations, which were then validated with Anton 2 at PSC.”
The supercomputer-generated simulations run by Bharat Poudel, first author on the study, specifically showed the effect of physical stimuli such as membrane tension, bilayer thickness and lipid chemistry on the structural properties of the AT1 receptor. Vanegas said that the simulations allowed the team to effectively understand mechanical activation at the molecular level.
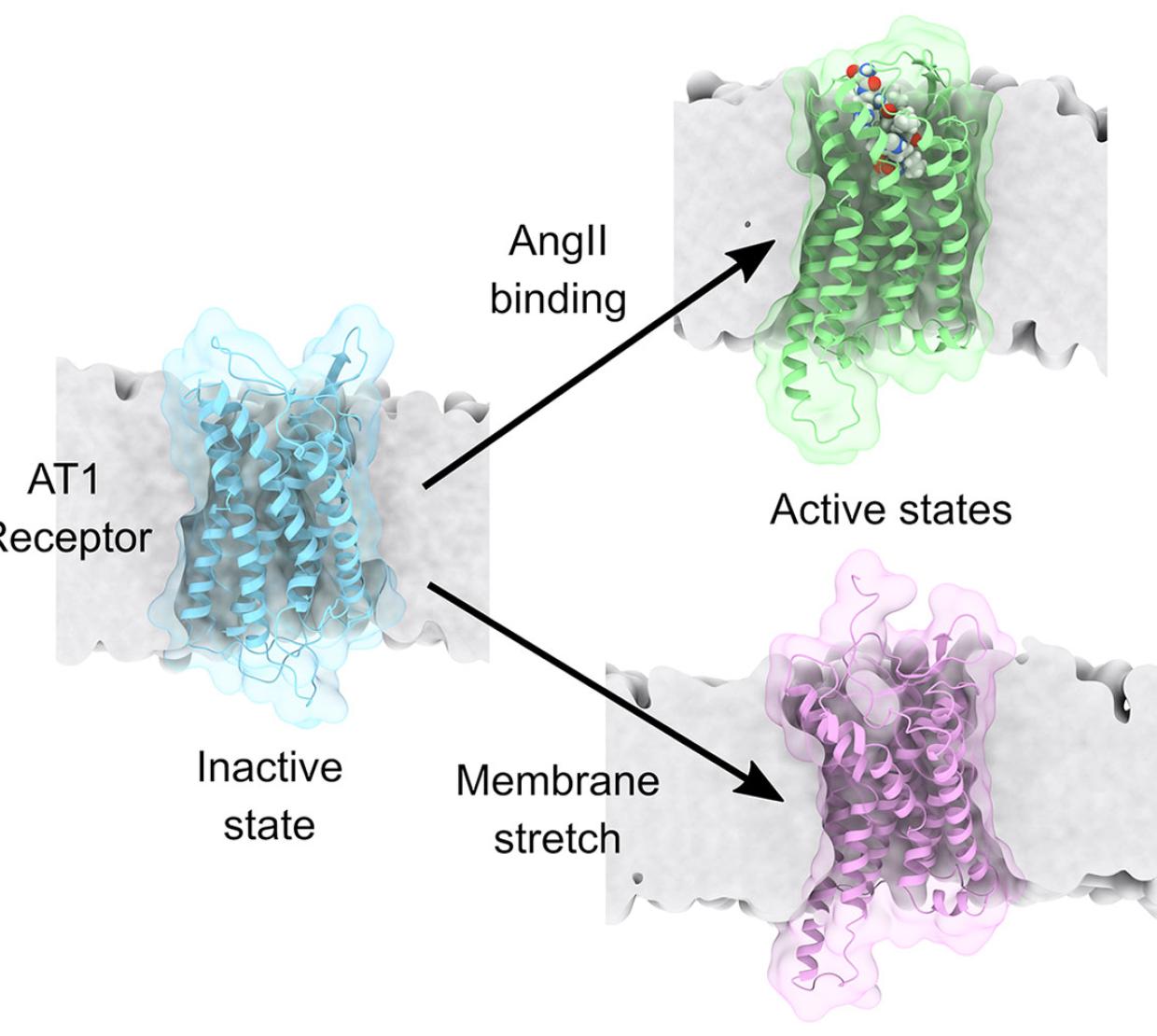
Applying membrane tension to the AT1 receptor in these simulations is analogous to tugging the edges of a cell and stretching it as if it was a rubber balloon. The stretching thins the membrane and has been discovered to change the stability of the active and inactive states of the protein.
“Our simulations showed that stretching the membrane stabilizes the AT1 receptor in a structural conformation similar to the active state induced by the chemical ligand angiotensin II, although it has some unique features that are different,” Vanegas said. “This is important because it is physiologically known that the AT1 receptor activates different signaling pathways with various functional outcomes depending on whether it is interacting with angiotensin II or if it’s activated by membrane stretch.”
While structural techniques such as X-ray crystallography and Cryo-EM have provided important structural details of activation/inactivation by ligands and drugs, it is difficult for these techniques to capture the effect of physical stimuli such as membrane tension because the protein structure cannot be readily frozen under the effect of those conditions. Yet Vanegas’ team was able to use the supercomputers to illustrate an unusual phenomenon.
“GPCRs are in general quintessential examples of the ‘lock and key’ model in biochemistry, where a special key – an agonist molecule such as angiotensin – is needed to activate the particular lock – the receptor such as AT1,” he said. “Within this context, activation by an external mechanical stimulus, which is chemically non-specific, is analogous to opening the lock by turning it with a screwdriver.”
Vanegas explained that the mechanical activation of the AT1 receptor works because the properties of the surrounding lipid membrane are intimately connected to its mechanical properties, such as membrane thickness and stiffness, and therefore external forces acting on the membrane change the way that the membrane interacts with the protein and drive particular molecular interactions to take place.
“The function of the receptor is extremely complex and used in numerous cellular signaling pathways,” he said. “Researchers like me look to discover more about how physical stimuli receptor activation can affect other cellular partners, as well as using this GPCR stimulation to understand the function of other mechanically activated proteins.”
Researchers on Vanegas’ team included Poudel, a recent graduate of the Materials Science graduate program at the University of Vermont, and Rajitha Rajeshwar, a former University of Vermont post-doctoral researcher who now works at Oak Ridge National Laboratory.
This research was supported by Extreme Engineering and Science Discovery Environment (XSEDE) resources at SDSC (allocation number TG-BIO210110). Time on Anton 2, which is hosted at PSC thanks to manufacturer D. E. Shaw Research, was provided through the National Institutes of Health (grant no. R01GM116961). The Vanegas lab is funded by the National Science Foundation CAREER program (award number CHE-1944892/2326678).
Media Contacts
Kimberly Mann Bruch
SDSC Communications